4



If you haven't already uploaded you upload here for free
https://www.impute.me/imputeme/
then go to the ethnicity module and put your ID number in
https://www.impute.me/ethnicity/
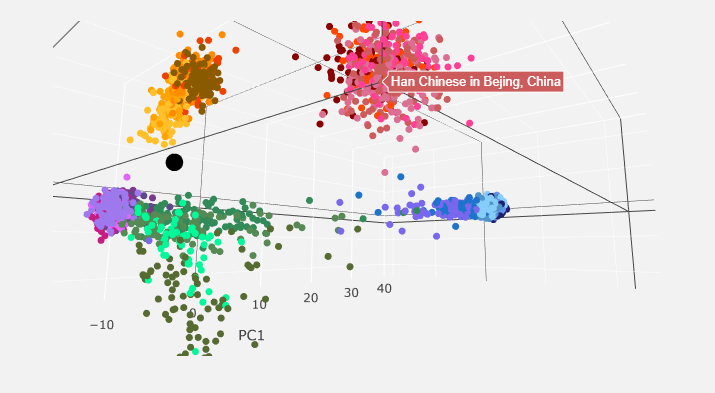
You get left with a 3D interactive PCA chart that maps your genome as a black dot in comparison to other populations. Here are some screenshots of mine














| Thumbs Up |
| Received: 2,549 Given: 1,670 |

2-6 days awaiting time after upload



| Thumbs Up |
| Received: 3 Given: 2 |

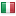
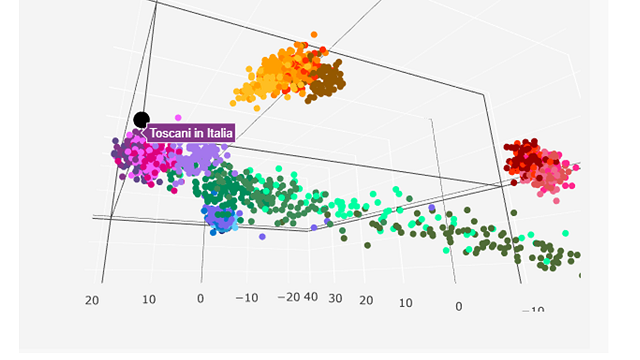
Here's mine - right next to another italian














| Thumbs Up |
| Received: 2,549 Given: 1,670 |














| Thumbs Up |
| Received: 2,549 Given: 1,670 |

https://www.tapatalk.com/groups/anth...80372-s15.htmlOriginally Posted by lassefolkersen

BUMB

Bump if anyone still hasn’t had a chance to upload
There are currently 1 users browsing this thread. (0 members and 1 guests)
 Latin America
Latin America
Bookmarks