0















| Thumbs Up |
| Received: 3,234 Given: 978 |














| Thumbs Up |
| Received: 13,747 Given: 3,217 |

Not so specific in my case:
But my version of 23andme didn't test for DF27.Most specific position on the YFull tree is R-P312
















| Thumbs Up |
| Received: 11,836 Given: 7,303 |

I have just learned that I'm negative for L332 and L329. This was the final crossroad before to see my cluster.
Greetings to fatherland Altai!














| Thumbs Up |
| Received: 3,337 Given: 3,930 |

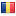
I guess I have just CTS3402.
















| Thumbs Up |
| Received: 1,541 Given: 1,154 |

ydna.JPG
y-dna2.JPG
So i have the continental "version" of the I2a2 haplogroup I-L701 (AKA I-cts10100), but how can find my sub-clade?















| Thumbs Up |
| Received: 5,083 Given: 2,784 |














| Thumbs Up |
| Received: 6,245 Given: 1,444 |













| Thumbs Up |
| Received: 3,234 Given: 978 |














| Thumbs Up |
| Received: 6,245 Given: 1,444 |

I thought of that already but I can't login into my old dna.land account which converted my ancestrydna file to 23andme for free.Do you know of any unix/linux scripts that will convert AncestryDNA to 23andme format ? I can't find any but perl or sed/awk with bash , in linux, would be the perfect tool for this kind of stuff man.
Maybe, I'll do it tomorrow if I remember. I think I have my ancestrydna raw data file converted to 23andme format on my old computer downstairs probably.Code:Error snps= cladeFinder returned: {"error":"unable to determine clade due to no positive SNPs"}













| Thumbs Up |
| Received: 3,234 Given: 978 |

This program does the job:
http://dnagenics.com/download/DnaKitStudiov28.zip
DNA KIT STUDIO
There are currently 1 users browsing this thread. (0 members and 1 guests)
Bookmarks