0

















| Thumbs Up |
| Received: 7,331 Given: 2,699 |

The crucial thing is to avoid using low coverage samples because the SNP overlap is always defined by the weakest link of the chain, and always use allsnps=YES (maxmiss=1 in Admixtools2, i believe). There is not much secret in choosing the pright, choose populations that don't violate the qpAdm assumption of no geneflow from pleft into pright (in practice, select prehistoric samples - the older the better), and make sure the populations in pright are all asymmetrically related to the populations in pleft.


















| Thumbs Up |
| Received: 1,249 Given: 524 |

You get "A" for effort. Nice heatmap.
They had some errors in the code. There's a new version dated yesterday of Admixtools. Remove it and re-install it. Also :
indicates you have too many uninformative non-polymorphic SNPs. To remove them set maxmaf=0.45593124 SNPs remain after filtering. 414780 are polymorphic
For FST it's crucial to have maxmiss at default which is 0 because you want all your samples to overlap each other exactly for unbiased results


















| Thumbs Up |
| Received: 1,249 Given: 524 |

The good thing about having SE and p-values is you get an idea which models are feasible unlike with Admixture or G25 where you get a bunch of feasible models and you have no idea which ones are no good (distance in G25 is NOT a substitute for p-values and SE by any means). Of course there are other issues besides model viability in G25.
For FST it's crucial to have maxmiss at default which is 0 because you want all your samples to overlap each other exactly for unbiased results
They had some errors in the code. There's a new version dated yesterday of Admixtools. Remove it and re-install it and re-run extract at default (maxmiss=0). When you run extract add the option maxmaf=0.45 to get rid of minor allele which have a frequency > 45% across all pops (very common old alleles shared by most global pops). Now I'm getting 940,000 polymorphic SNPs on the Simons samples at maxmiss=0 with the latest download of Admixtools.
Also Dilawer informed me that Plink doesn't maintain allele order. This can screw calculations up in Admixtools. Here is a couple of rows from the .snp file
rs199706086 1 0 10250 A C
rs112750067 1 0 10327 T C
rs201725126 1 0 13116 G T
rs200579949 1 0 13118 G A
rs180734498 1 0 13302 T C
rs79585140 1 0 14907 G A
rs75454623 1 0 14930 A G
Column 5 is supposed to be for the Reference or Ancestral allele and col 6 for the Derived or Minor allele. If you check dbSNP website you'll notice that the ones I bolded are wrong order. In other words one should be T G and the other A G. Plink screws up allele order.
I fixed the allele order in my .snp file using Dilawer's script so now it agrees with dbSNP. There were thousands of such mistakes in my .snp file














| Thumbs Up |
| Received: 4,863 Given: 2,946 |

This prints all columns and all rows of tables, prints only 3 significant digits, and doesn't display negative numbers in red (https://tibble.tidyverse.org/reference/formatting.html): `options(tibble.width=Inf,tibble.print_max=Inf, pillar.sigfig=3,pillar.neg=F)`.
`options(width=Sys.getenv("COLUMNS"))` or `options(width=180)` increases the width of the terminal.
`print(tbl,width=Inf,n=Inf)` or `as.data.frame(tbl)` displays a whole tibble. This displays an HTML table in a browser: `install.packages("formattable");library(formattable);formattable(tbl)`.
This removes columns from a tibble and formats the table as CSV where doubles have 3 digits after the decimal point:
This omits models with only one population (where f4rank is 0) and models that are not feasible (with one or more negative weight). It then sorts the remaining models by their p value:Code:> qp$popdrop%>select(!c(pat,wt,dof,dofdiff,chisqdiff,p_nested))%>%mutate(across(where(is.double),round,3))%>%format_csv%>%cat chisq,p,f4rank,Nganasan,Norway_N_HG.SG,Russia_AfontovaGora3,Turkey_Epipaleolithic,feasible,best 3.251,0.354,3,0.128,0.132,0.116,0.625,TRUE,NA 15.538,0.004,2,0.151,1.265,-0.416,NA,FALSE,TRUE 5.152,0.272,2,0.144,0.302,NA,0.554,TRUE,TRUE 3.827,0.43,2,0.123,NA,0.176,0.701,TRUE,TRUE 11.855,0.018,2,NA,-0.814,0.64,1.173,FALSE,TRUE 28.572,0,1,0.072,0.928,NA,NA,TRUE,NA 116.62,0,1,-0.108,NA,1.108,NA,FALSE,NA 11.9,0.036,1,0.179,NA,NA,0.821,TRUE,NA 22.973,0,1,NA,1.388,-0.388,NA,FALSE,NA 28.13,0,1,NA,0.622,NA,0.378,TRUE,NA 16.715,0.005,1,NA,NA,0.272,0.728,TRUE,NA 1386.434,0,0,1,NA,NA,NA,TRUE,NA 32.704,0,0,NA,1,NA,NA,TRUE,NA 125.18,0,0,NA,NA,1,NA,TRUE,NA 36.405,0,0,NA,NA,NA,1,TRUE,NA
This saves the popdrop table to a CSV file (I know my pright sucks or whatever):Code:> qp$popdrop%>%dplyr::filter(feasible==T&f4rank!=0)%>%arrange(desc(p))%>%dplyr::select(!c(pat,wt,dof,chisq,f4rank,feasible,best,dofdiff,chisqdiff,p_nested))%>%mutate(across(where(is.double),round,3))%>%as.data.frame p Nganasan Norway_N_HG.SG Russia_AfontovaGora3 Turkey_Epipaleolithic 1 0.430 0.123 NA 0.176 0.701 2 0.354 0.128 0.132 0.116 0.625 3 0.272 0.144 0.302 NA 0.554 4 0.036 0.179 NA NA 0.821 5 0.005 NA NA 0.272 0.728 6 0.000 NA 0.622 NA 0.378 7 0.000 0.072 0.928 NA NA
This generates a stacked bar chart of the models sorted by their p value:Code:target="Finnish" left=c("Turkey_Boncuklu_N.SG","Latvia_HG","Norway_N_HG.SG","Russia_HG_Karelia","Russia_HG_Tyumen","Russia_AfontovaGora3","Nganasan") right=c("Mbuti.DG","Mixe.DG","Ami.DG","Czech_Vestonice16","Papuan.DG","Ethiopia_4500BP_published.SG","Russia_Kostenki14","Ju_hoan_North.SDG","Morocco_Iberomaurusian") pops=c(left,right,target) unlink("f2",recursive=T) extract_f2(pref="g/v44.3_HO_public/v44.3_HO_public",pops=pops,outdir="f2") f2=f2_from_precomp("f2") qp=qpadm(f2,left=left,right=right,target=target) qp2=qp$popdrop%>%dplyr::filter(feasible==T&f4rank!=0)%>%arrange(desc(p))%>%dplyr::select(!c(wt,dof,chisq,f4rank,feasible,best,dofdiff,chisqdiff,p_nested)) write_csv(qp2,"/tmp/a")
Code:library(tidyverse) library(cowplot) library(reshape2) t=read_csv("/tmp/a") abbr=c("Turk","Latv","Norw","Kare","Tyum","AG3","Ngan") l=lapply(t$pat,function(x)abbr[unlist(gregexpr("0",x))]%>%paste(collapse=" ")) t$lab=paste0(l," (",sub("^0","",sprintf("%.3f",t$p)),")") t=t[-c(1,2)] t2=melt(t,id.var="lab") p=ggplot(t2,aes(x=fct_rev(factor(lab,level=t$lab)),y=value,fill=variable),label=pvalue)+ geom_bar(stat="identity",width=1,position=position_fill(reverse=T))+ geom_text(aes(label=round(100*value)),position=position_stack(vjust=.5,reverse=T),size=4)+ coord_flip()+ theme( axis.text.x=element_blank(), axis.text=element_text(color="black"), axis.ticks=element_blank(), axis.title.x=element_blank(), legend.box.just="center", legend.box.margin=margin(0,unit="cm"), legend.box.spacing=unit(.05,"in"), legend.direction="horizontal", legend.justification="center", legend.margin=margin(0,unit="cm"), legend.text=element_text(size=12), legend.title=element_blank(), panel.border=element_blank(), text=element_text(size=18) )+ guides(fill=guide_legend(ncol=3,label.position="right",byrow=T))+ scale_x_discrete(expand=c(0,0))+ scale_y_discrete(expand=c(0,0))+ xlab("")+ scale_fill_manual("legend",values=c("#be661f","#66f6ff","#3397f5","#22419c","#39de39","#157f0a","#ef50ed")) ggdraw(p) leg=get_legend(p) p=p+theme(legend.position="none") ggdraw(plot_grid(p,leg,ncol=1,rel_heights=c(1,.2))) ggsave("output.png",width=7,height=7)
Why do Finns get such a high percentage of Turkey_Boncuklu? Is it because of bad right populations or something?
Last edited by Komintasavalta; 03-08-2021 at 05:11 PM.
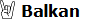















| Thumbs Up |
| Received: 1,541 Given: 1,154 |

Hi everyone, could someone run with qpAdm North_Italians, Tuscans and Sicilians with this model?
WHG
Steppe_EMBA (Samara i think it's ok)
Barcin_N
Iran_N
Levant_PPNB
I want to see if Levant_PPNB it's necessary to run italians, in particulary southerns.
Thank you.













| Thumbs Up |
| Received: 3,471 Given: 1,541 |

which of these should be used as outgroups when modelling modern pops wiith mesolithic/neolithic samples?
Austria_Krems1_1 I2483 Austria_Krems1_2_twin.I2483 I2484 Austria_Krems1_2_twin.I2483_all I2484_all Austria_KremsWA3 I1577 Belgium_UP_GoyetQ116_1_published GoyetQ116-1_udg_published Belgium_UP_GoyetQ116_1_published_all GoyetQ116-1_published Belgium_UP_GoyetQ376-19_published GoyetQ376-19_published_d Belgium_UP_GoyetQ53_1_published_lc GoyetQ53-1_published_d Belgium_UP_GoyetQ56_16_published_lc GoyetQ56-16_published_d Belgium_UP_Magdalenian GoyetQ-2 Belgium_UP_Magdalenian_udg GoyetQ-2_udg China_Tianyuan Tianyuan Czech_Pavlov1 Pavlov1_d Czech_Vestonice13 Vestonice13_d Czech_Vestonice14_lc Vestonice14_d Czech_Vestonice15 Vestonice15_d Czech_Vestonice16 Vestonice16 Czech_Vestonice43 Vestonice43_d France_Rigney1_published Rigney1_published_d Germany_Brillenhohle_published_lc Brillenhohle_published_d Germany_Burkhardtshohle_published Burkhardtshohle_published_d Germany_HohleFels49_published HohleFels49_published_d Germany_HohleFels79_published_lc HohleFels79_published_d Italy_South_HG_Ostuni1 Ostuni1_d Italy_South_HG_Ostuni2 Ostuni2_d Italy_South_HG_Paglicci108_published_lc Paglicci108_published_d Italy_South_HG_Paglicci133_published Paglicci133_published Romania_Cioclovina_published_lc Cioclovina1_published_d Romania_Muierii Muierii2_d Romania_Oase Oase1_d Russia_Kostenki12 Kostenki12 Russia_Kostenki14 Kostenki14 Russia_Kostenki14.SG Kostenki14.SG Russia_Sunghir1.SG Sunghir1.SG Russia_Sunghir2.SG Sunghir2.SG Russia_Sunghir3.SG Sunghir3.SG Russia_Sunghir4.SG Sunghir4.SG Russia_Ust_Ishim_HG_published.DG Ust_Ishim_published.DG Russia_Ust_Ishim.DG UstIshim_snpAD.DG Russia_Yana_UP.SG Yana_old.SG Russia_Yana_UP.SG Yana_old2.SG Spain_ElMiron ElMiron_d
Kostenki14 and Ust-Ishim were used here, but this was a couple of years ago and new samples have been published since, so should something be added?
https://eurogenes.blogspot.com/2017/...lithic-to.html













| Thumbs Up |
| Received: 3,471 Given: 1,541 |

most of these Paleolithic samples were analyzed here:
https://www.ncbi.nlm.nih.gov/pmc/art...ort=objectonly
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4943878/
I ran a PCA on all Reich samples older than 8000 ybp, to see into which clusters the more recently published samples fall. I removed Neanderthals and Denisovans, and African, Middle Eastern and Amerindian samples because they were outliers and skewed the European dimensions. I didn't do ld and maf pruning because they PCA. I guess Paleolithic samples probably require completely different ld and maf settings from modern pops.
The result is very similar to the MDS plot from the study:
The conclusions would be Sunghir samples fall into the Vestonice cluster, Yana clusters with Ust-Ishim, and Geometric and Azilian samples plot with El Miron.
















| Thumbs Up |
| Received: 11,836 Given: 7,303 |

I was trying to extract new Turkish samples but received a few errors during the process. Does the files needs to be imputed in Linux by referencing to their genetic linkage?(by using SHAPEIT perhaps) At least that's what I understood, although have no idea how to do it.
Any tips on what is going on here?
Files can be found here in case someone would like to try it.
Code:+ extract_f2("originknownturkish",outdir = "balkanturks", pops = pops) i Reading allele frequencies from PLINK files... Warning: 1 parsing failure. row col expected actual file 1460 X6 embedded null 'originknownturkish.fam' i originknownturkish.geno has 1460 samples and 423261 SNPs i Calculating allele frequencies from 1 samples in 1 populations i Expected size of allele frequency data: 102 MB 423k SNPs read... √ 423261 SNPs read in total ! 421395 SNPs remain after filtering. 0 are polymorphic. i Allele frequency matrix for 421395 SNPs and 1 populations is 34 MB i Computing pairwise f2 for all SNPs and population pairs requires 67 MB RAM without splitting i Computing without splitting since 67 < 8000 (maxmem)... Error in cpp_get_block_lengths(numchr, dat[[distcol]], blgsize) : upper value must be greater than lower value Ek olarak: Warning messages: 1: In get_block_lengths(afdat$snpfile, blgsize = blgsize) : No genetic linkage map or base positions found!Each chromosome will be its own block, which can make standard error estimates inaccurate. 2: In get_block_lengths(afdat$snpfile[poly, ], blgsize = blgsize) : No genetic linkage map found! Defining blocks by base pair distance of 2e+06
There are currently 1 users browsing this thread. (0 members and 1 guests)
Bookmarks